Our laboratory focuses on understanding structure-function relationships in nucleic acid systems. In addition to NMR spectroscopy, we use biochemical and other biophysical methods (SAXS, single molecule FRET, x-ray crystallography) to investigate the kinetic, motional, and structural properties of the component molecules.
The data obtained from these methods complement physiological and genetic information. One focus of the lab centers on post-transcriptional modification of tRNA and the contributions of modifications to tRNA-mediated gene expression through T-box riboswitch mechanism. The bases of tRNA molecules are modified in all cells. Modification, especially within the anti-codon loop, has been linked to translational fidelity and efficiency and regulation in prokaryotes. We are examining structural properties and the kinetic parameters of T-box riboswitch-tRNA binding to develop mechanistic models. Modifications throught tRNA affect the molecule's structure and stability. We are elucidating the roles of various tRNA modifications on the efficiency of riboswitch function in the cell and on the kinetics of riboswitch-tRNA binding. These studies also will extend our understanding of the contributions of tRNA modification to Gram positive bacterial cell physiology and fitness.
Non-coding RNAs (nc-RNA), including rRNAs, riboswitches, and pre-microRNAs, often have an architecture containing a variety of non-canonical features that are key for the molecules' structures and function, therefore, the detailed structure provides an important backdrop to interpret functional and mechanistic studies. Small molecules that bind to specific RNA structure motifs can become powerful tools for probing RNA conformation and folding and to facilitate studies of specific RNA activities in vitro and in vivo. We are using NMR spectroscopy to define the breadth of conformational space accessed by a set of frequently occurring non-canonical RNA motifs. A computational workflow, that integrates the NMR-defined structure information and in silico screening of virtual chemical libraries is being used to identify chemical compounds that selectively bind to these non-canonical motifs. The interactions of these small molecules with RNAs are being characterized using biophysical and high-resolution structure methods including NMR spectroscopy and X-ray crystallography.
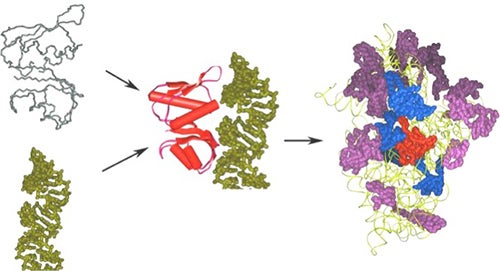
Another system involves the interaction of primary ribosomal proteins with their RNA targets and provides the opportunity to not only explore the specificity of RNA-ligand interactions, but also to examine the co-folding of protein and RNA molecules and maturation of the 30S (prokaryotic) and 40S (eukaryotic) ribosomal subunit. The primary RNA binding site for the S8 protein folds around a Mg2+ ion to form an unusual tertiary structure made up of non-Watson-Crick interactions. Once bound, S8 stabilizes the interaction of other ribosomal proteins with the 16S rRNA and guides folding of the RNP subdomains to yield the mature 30S particle.

